UPTIDER, a highly collaborative project requiring a robust OSE
UPTIDER is a highly collaborative programme intended to enhance our understanding of metastatic breast cancer32. Figure 1 and Supplementary Fig. S1 describe the landmark events within UPTIDER highlighting both the required intra-team collaboration and the importance of providing the team with prompt access to latest information. This also explains the logistical and scientific need for developing and maintaining an OSE in UPTIDER (Fig. 2 and Methods). In this regard, as of May 2025, the UPTIDER OSE helped facilitate the acquisition and annotation of >15000 samples from 39/45 enroled patients. These samples were acquired from >30 sites of solid tissue and 7 distinct sources of liquid biopsy leading to a median of 300 samples per autopsy. Figure 3 provides a global overview of the components of the OSE and their trajectory based on an established framework design and timeline, illustrating the foundation required to implement open science practices within UPTIDER. Complementing Fig. 3, Supplementary Fig. S2 presents the architectur diagram of the OSE highlighting data and code flows between user, storage, and computational resources.
A flowchart and timeline describe the key processes within UPTIDER attributed to each of the players in the team from patient inclusion to study publication. The flowchart elucidates the degree of inter-player communication and collaboration within the UPTIDER team, emphasizing the logistical and scientific need for a UPTIDER OSE to support its research objectives. The timeline outlines the major events between patient inclusion and study publication. Rectangles in pink describe the patient trajectory within the OSE, purple describes the actions performed by the entire team, followed by actions of the researcher, clinician, pathologist, bioinformatician and lab technician in orange, yellow, light green, blue and light blue rectangles, respectively. The role of a researcher can be embodied by a clinician, lab technician, pathologist, scientist and/or research data manager. Solid arrows indicate dependent relationships between actions, where the previous action must end before the next one can start. Dashed lines indicate a parallel flow to the main action. Not displayed on the figure, but during tissue donation, samples are also processed for external institutional collaborators. Additionally, data and material sharing are facilitated after approval of the ethics committee and the signing of material and data transfer agreements. These agreements are drafted in collaboration with the KU Leuven legal department. For access to raw sequencing data, a request needs to be made to the data access committee via the EGA portal. Extended version of the flowchart is available in Supplementary Fig. S1. eCRF: electronic case report form, LIMS: lab information management system, OSE: open science environment, UPTIDER: UZ/KU Leuven Post-mortem Tissue Donation to Enhance Research. Created with Microsoft Visio.
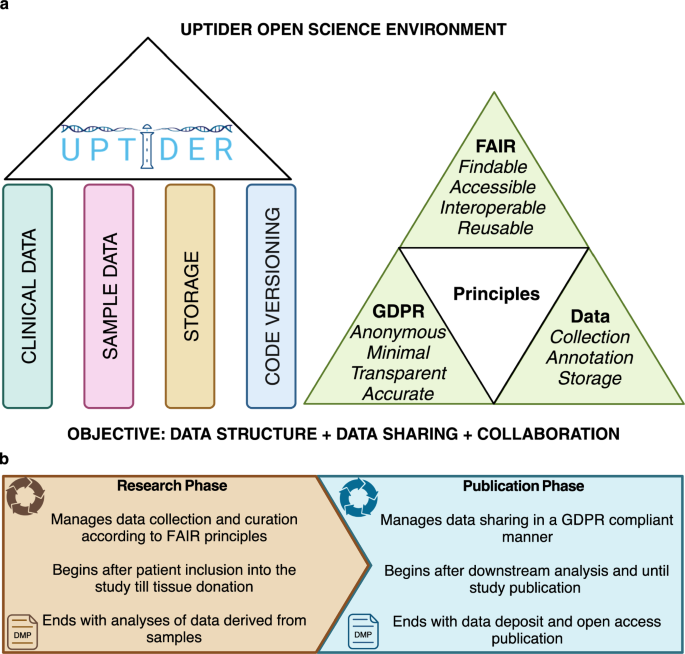
a Graphical abstract describing the 4 main structural pillars used to design the components of the UPTIDER OSE, followed by the key guiding principles which were incorporated into the design. These enabled the OSE to satisfy the key objective of the UPTIDER OSE, which was to provide a robust data structure during data collection, annotation and storage while making data sharing convenient and collaboration feasible. b Graphical abstract describing the two principal phases within the UPTIDER OSE during each research lifecycle. While these phases are linked by a linear flow from one to the other, each of these phases were also independently governed and supported by the overarching DMP. This created an independent provision for continuous development and improvement to both phases throughout the research lifecycle, making the OSE an adaptive entity. Created with biorender.com. DMP: data management plan, OSE: open science environment, UPTIDER: UZ/KU Leuven Programme for post-mortem Tissue Donation to Enhance Research.
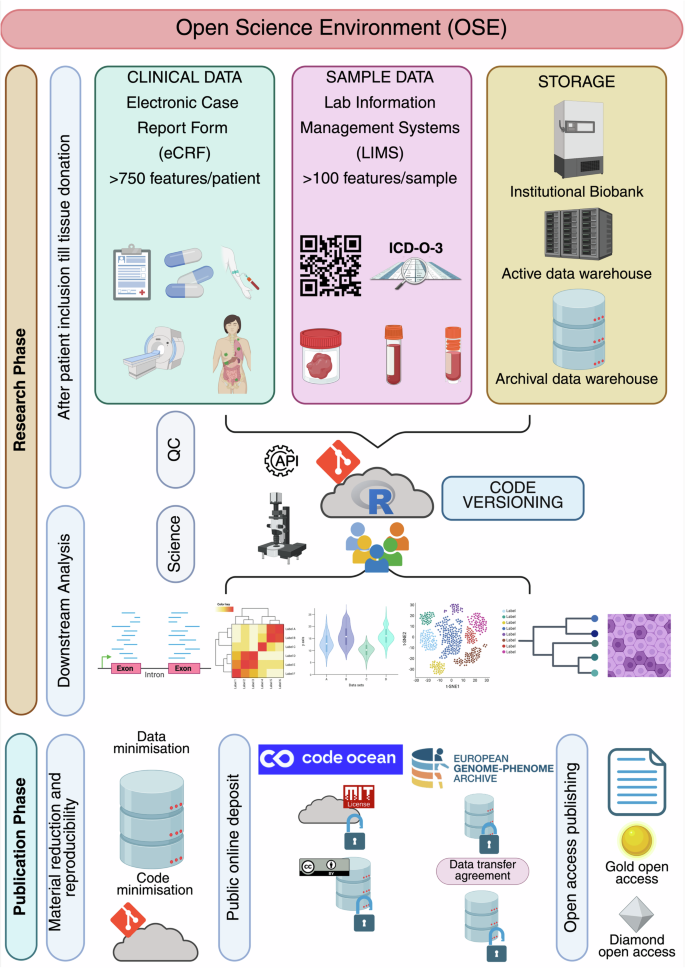
A description of the structural pillars and phases of the UPTIDER OSE along with their features. At the start of the research lifecycle, patients are included into UPTIDER. Clinical data, including more than 750 clinical features, are collected per patient using an eCRF in line with GDPR principles. Sample metadata, including more than 100 features per sample, are collected using a LIMS in line with FAIR principles. Sample data and sample generated data are stored in the institutional biobank and GDPR protected data warehouses that comply with regulatory frameworks. Following tissue donation, data curation and generation, quality checks and downstream analyses are performed for various UPTIDER sub-studies using the distributed code versioning system. Once research manuscript is sent for publication, material reduction (code and data) is performed in compliance with GDPR principles for data sharing and reproducibility, meaning that only the code and data essential to reproduce the study are shared. This material is then deposited into secure online public access environments such as Code Ocean under appropriate licensing for data re-use and reproducibility. The raw data is deposited into European Genome-Phenome Archive under restricted access, which can be requested following a data transfer agreement. Similarly sample access can be requested following a material transfer agreement. In parallel, the research manuscript undergoes peer-review and gets published in either a gold or a diamond open-access journal. GDPR: general data protection regulation, UPTIDER: UZ/KU Leuven Programme for post-mortem Tissue Donation to Enhance Research. Created with biorender.com.
Capturing clinical data
To improve clinical outcomes of future patients by translating current fundamental research into the clinic, up-to-date access to patient electronic health record (EHR) is needed to design research objectives and derive meaningful conclusions from patient data33,34. As such, patients with metastatic cancer are treated with multiple treatment regimens, undergo multiple blood draws and biopsies, as well as several surgeries and imaging examinations throughout their disease course. This means that each patient comes with a complex medical and clinical history, which needs to be combined with sample data to perform translational research. To satisfy this requirement and to enable protected access to data for the entire team, an electronic case record form (eCRF) needed to be designed. The first phase was to identify pivotal clinical features from the patient EHR for our study requirements and then develop a structure using a founder document (Fig. 3, Fig. 4 Phase I and Supplementary Table S1). This was an independent document that co-existed alongside the eCRF. The key purpose of this document was to capture proceedings from the brainstorming and design sessions for features that were to be included. After consensus was achieved on the feature structure from the founder document, an eCRF was registered. A second phase was then initiated to design and implement this structure into the eCRF (Fig. 4, Phase II). Constant collaboration with the clinical trial centre (CTC) at UZ Leuven was crucial in helping refine our requirements and feature structure before efficiently translating them into an eCRF. Once the eCRF was designed, internal quality checks were added to ensure accurate data input. We developed more than 25 internal quality checks (QC) by adding branching logic and data validation to specific fields, preventing inconsistent information. Instruments resembled a survey environment with structured drop-down menus and multiple-choice answers to ease data recording and to reduce missing data. Helper texts were added to increase consistent reporting. Predefined missing codes were implemented to capture reason of missing data to identify potential bias in downstream analyses. Additionally, free-text fields were minimized to reduce unstructured information. For features requiring recurrent updates such as team member and treatment regimens, structured query language (SQL) queries were developed to ease item addition into pre-existing dropdown lists. Once implemented, the eCRF was thoroughly tested by the entire team. Here, the founder document was instrumental in communicating with the UPTIDER team to capture feedback and track progress (Fig. 4, Supplementary Table S1). Post-validation, the eCRF was submitted for production (Fig. 4, phase III), allowing the team to record patient EHR data (Fig. 2). At this stage, modifications to the eCRF had to be as minimal as possible, since all modifications must be approved by the institution while ensuring no data loss. Eventually, more than 750 unique features per patient were recorded in the eCRF (Fig. 3) to capture clinical and autopsy data. The codebook of the UPTIDER eCRF can be found in the Supplementary Table S2. During data entry, each participating patient was encoded with a unique ID that was used to identify and annotate each feature in the patient record. The structure of the eCRF together with an application programming interface (API) allowed for data entry, export and reporting (Fig. 4, Phase III). The provision of role-based access and multifactor authentication (MFA) selectively enabled the team to securely record authorised clinical and non-clinical information in accordance with GDPR principles. Taken together, we have demonstrated the value of inter-team collaboration and highlight the extensive processes involved in the conception, development, testing, and production of the eCRF.
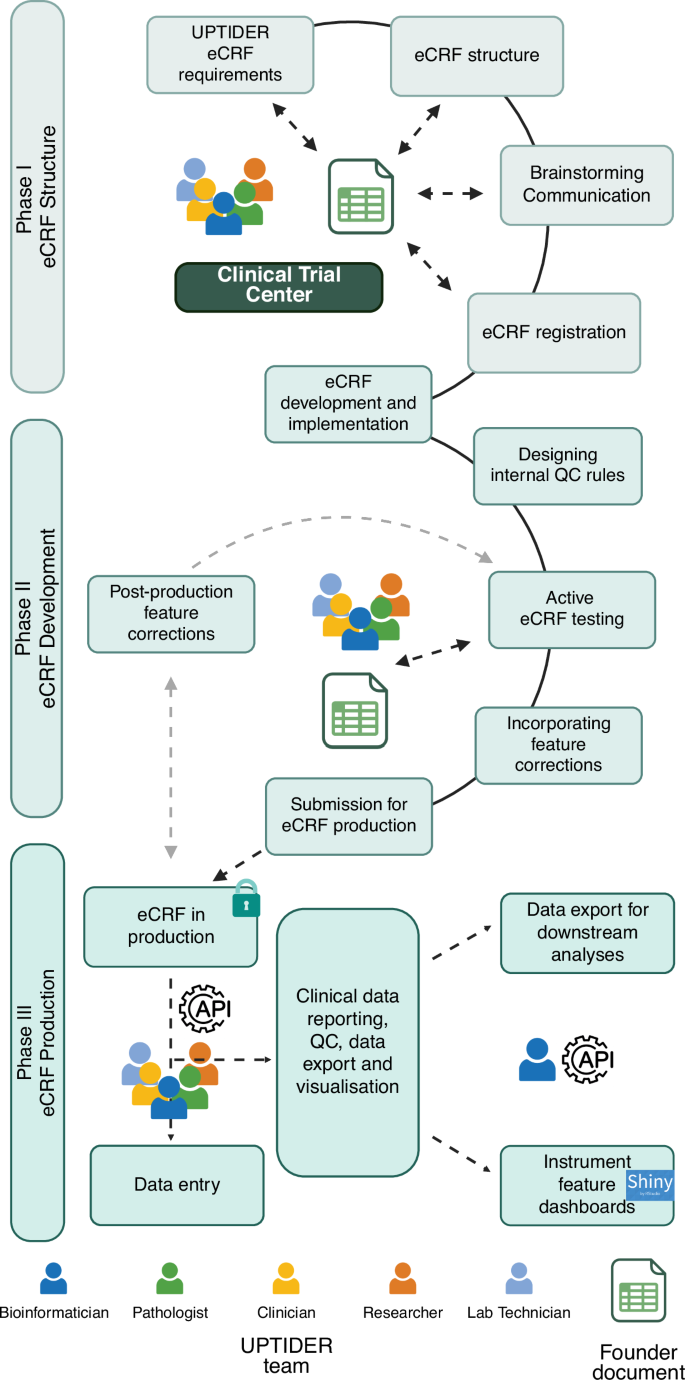
A brief description of the methodology employed to design and implement the UPTIDER eCRF as a sequential process. The methodology involved 3 different phases linked to the eCRF structure, development and production and ultimately the final design had to comply with the GDPR principles. Constant communication between the UPTIDER team, and the clinical trial centre facilitated the development of the founder document and the preliminary registration of the eCRF emphasized by the double-tailed arrows. During development, internal QC rules and clinical features were designed, tested and implemented using the founder document in communication with the UPTIDER team. Once the eCRF was submitted for production, post-production feature corrections were kept to a minimum, highlighted by the grey-dashed arrows, to prevent data corruption and loss during data entry. Data entry was facilitated by both manual entry with the team and with SQL query via API. This API also helped in the data export of the eCRF to be used for data curation, visualization and reporting. API: application programming interface, eCRF: electronic case report form, GDPR: general data protection regulation, SQL: structured query language, UPTIDER: UZ/KU Leuven Programme for post-mortem Tissue Donation to Enhance Research. Created with biorender.com. Schematic based on REDCAP (v15.0.37).
Recording sample data and metadata
An UPTIDER autopsy can yield up to 600 solid and liquid samples32. This necessitates a robust data and metadata structure, that can be routinely updated by the team after using various research techniques on these samples (Fig. 1). We therefore customised our lab information management system (LIMS) by initiating an appropriate feature structure design (Fig. 5, Phase I). The structure needed to accommodate the legal requirement for the use of human body materials (HBM), standard preanalytical coding for biospecimens (SPREC) metadata reporting standards and institutional biobank recommendations (Supplementary Table S3), while also satisfying the internal metadata needs of UPTIDER. At this stage, collaboration and correspondence with the biobank was crucial. Like the eCRF, this structuring phase was supported by a founder document (Supplementary Table S3), which registered metadata design sessions and intra-team communication. In UPTIDER, wherever possible, a tissue site is sampled under multiple conditions in mirror (formalin-fixed paraffin embedded (FFPE), Fresh frozen in optimal cut temperature medium (FF-OCT) and fresh frozen (FF) for instance) for various analyses (Fig. 3). FFPE allows histopathological assessment for essential tumour characterization, while FF enables next-generation sequencing (NGS) of DNA and/or RNA among others. Therefore, information from FFPE samples must be linked to FF samples used for sequencing, which lab technicians extract and bioinformaticians analyze. Thus, mirrored sample linkage was a mandatory requirement within the LIMS. Another fundamental requirement was the generation and linkage of derived samples such as DNA libraries that originate from a parent sample (FFPE, FF-OCT or FF). Selecting a suitable LIMS to fulfil these requirements while being financially sustainable was crucial. As an institutional LIMS was not available, we explored different players in the market. Post-identification and feature approval, we initiated the LIMS development using the founder document (Fig. 5, Phase II). At this stage, we added approved metadata feature components into the LIMS and included a unique serial ID for each patient, complementing the LIMS identifier. This created an internal double authentication system, securing sample identification and tracking. Internal QCs were also designed to prevent incorrect data entry (Fig. 5). Additionally, active testing of the LIMS was facilitated by the founder document with documentation of feature corrections from the team along with a status tracker. We then initiated the final phase of the LIMS and submitted our LIMS for production (Fig. 5, Phase III). Here, data entry was largely supported by automated code scripts. As a result, more than 100 features per sample have been implemented including sample label, date of collection and organ code according to the ICD-O-3 organ classification35 (Supplementary Fig. S3). Mirrored and derived samples are automatically handled with a R script using the API. An audit trail logs user activity across samples. Each sample is given a label with a QR code and structured text to guide sample collection (Supplementary Fig. S3). This QR code is linked to the LIMS ID and helps keep track of the sample throughout its lifecycle. Additionally, to avoid data loss, samples registered but not collected are never deleted but rather, archived. A version of the LIMS codebook can be found in Supplementary Table S4. The LIMS and the eCRF comply with FAIR principles and are linked by the patient ID. Electronic lab notebooks (ELN) are used to document and track protocols for the sample processing. Each notebook is assigned to a specific protocol with version numbers and a timeline to trace the eventual development of the standard operating protocol (SOP)32. Taking these into consideration, we report on the collaborative efforts on designing the appropriate methodology for our custom metadata structures and selecting a LIMS that could accommodate legal requirements and internal needs while ensuring accurate sample tracking.
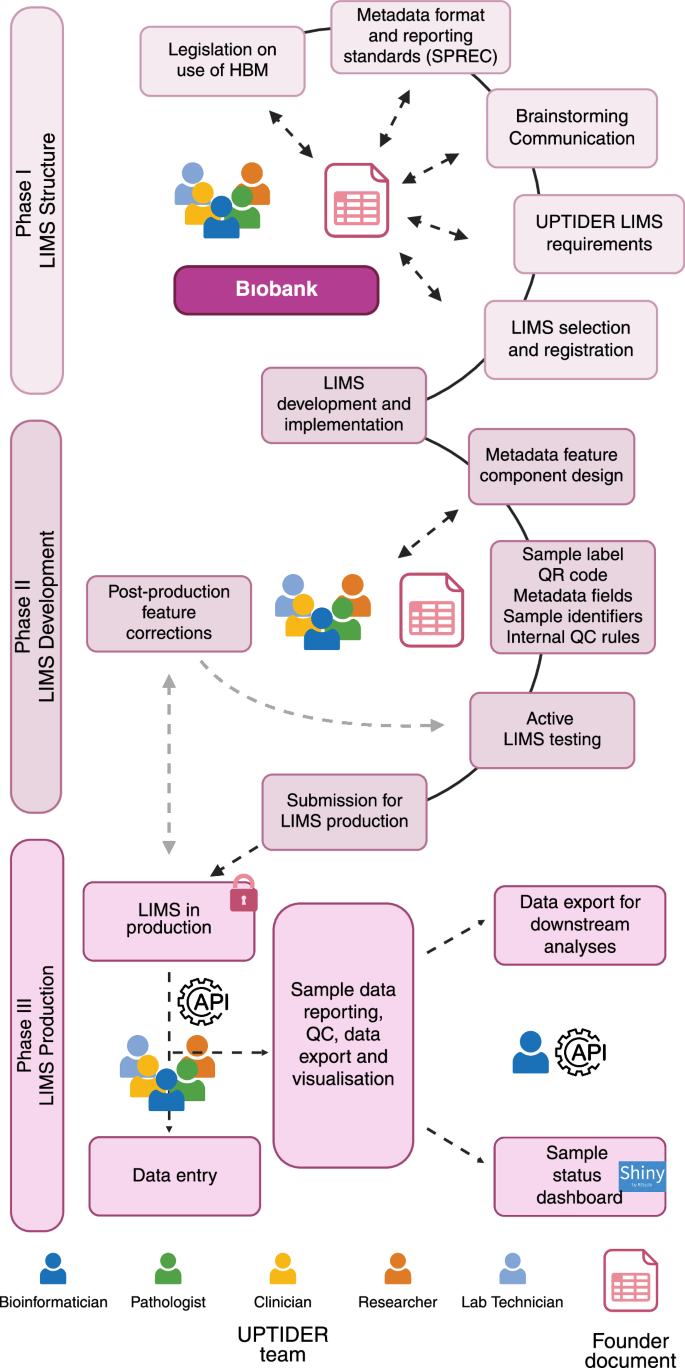
A brief description of the methodology employed to design and implement the UPTIDER LIMS as a sequential process. The methodology involved 3 different phases linked to the LIMS structure, development and production and the final design had to comply with the regulatory legislation, SPREC reporting standards for metadata and FAIR principles. Constant communication between the UPTIDER team, and the institutional biobank facilitated the development of the founder document and the selection of the LIMS emphasized by the double-tailed arrows. During development, internal QC rules and metadata features were designed, tested and implemented using the founder document in communication with the UPTIDER team. Once the LIMS was submitted for production, post-production feature corrections were kept to a minimum, highlighted by the grey-dashed arrows, to prevent data corruption and loss during data entry. Data entry was facilitated by an SQL query via API. This API also helped in data export of the LIMS to be used for data curation, visualization and reporting. API: application programming interface, FAIR: findable, accessible, interoperable and reusable, LIMS: lab information management system, SPREC: standard preanalytical code, SQL: structured query language, UPTIDER: UZ/KU Leuven Programme for post-mortem Tissue Donation to Enhance Research. Created with biorender.com. Schematic based on Labcollector (v6.262).
Identifying sources for sample and data storage
For long-term usage and sharing of UPTIDER samples, appropriate storage and inventory solutions needed to comply with GDPR and FAIR principles while supporting automated data annotation. All samples collected during tissue donation have been securely stored at the biobank under restricted access. Our LIMS supports bio-tracking by registering sample storage locations and is continuously improved to maintain accuracy in metadata reporting. Additionally, the LIMS metadata structure aligns with biobank requirements, enabling automated metadata sharing and compatibility with initiatives like the new European Health Data Space for enhanced public health research36.
Raw data such as sequencing outputs and digitalized histopathological tissue sections, or processed data resulting from downstream analyses are stored in a GDPR protected archival data warehouse in distinct locations (Fig. 6a). This warehouse provides a secure environment with protected role-based access and MFA. Regular backups help prevent data loss. Currently, our data occupies approximately 35 terabytes of storage within the warehouse. For frequently accessed data, we use an active data warehouse to increase transfer speed, but it does not provide backups. Transfer between the storage solutions are achieved by Globus37 which enables secure data transfer and tracking in the background (Fig. 6a). The link between clinical data, sample data and raw data is collected in a live Excel 365 document (Supplementary Table S5). This document allows for concurrent access and ensures that each team member accesses the most up-to-date information regarding the samples. This file also gets a backup in a separate location for data recovery. Collectively, the LIMS, storage solutions, and the live document facilitate storage and access of samples and sample data, ensuring research integrity and transparency.
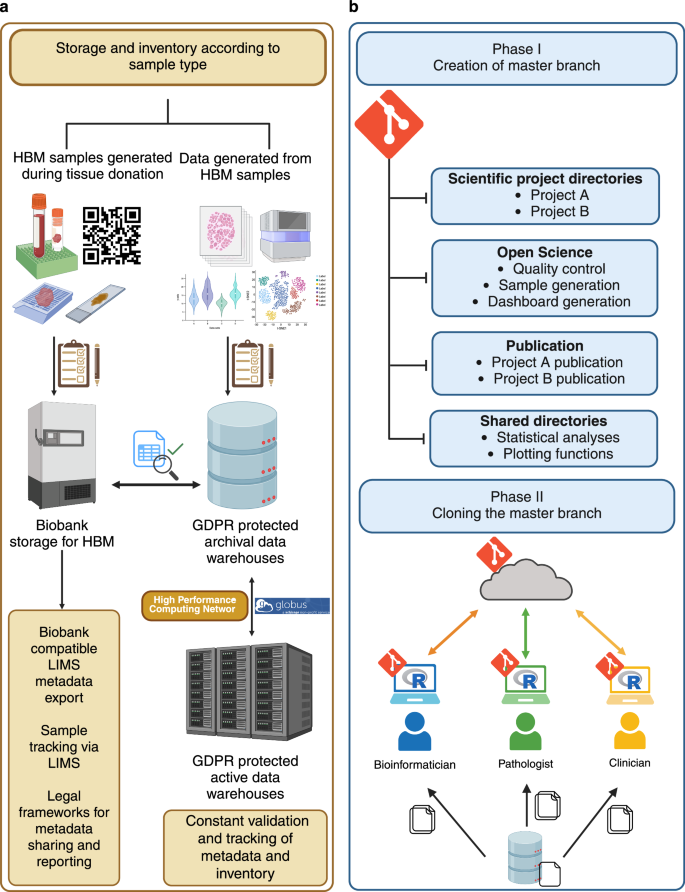
a Graphical description regarding the selection of storage solutions to store UPTIDER sample data and generated data. For samples collected after patient inclusion into UPTIDER, inventory is performed and stored at the institutional biobank with bio-tracking facilitated by the UPTIDER LIMS. The LIMS metadata structure is compatible with the regulatory requirements of the biobank for use of HBM, while making data sharing, depositing and retrieval convenient. For data and results generated from these samples with various research technologies, inventory is performed, and they are stored in GDPR protected archival data warehouses. A live excel 365 document is used to couple information from samples sent for NGS with their corresponding LIMS metadata. High-performance computing networks are used to transfer and process these sequencing files between the network and the archival warehouse. For frequently used data, active data warehouses and high-performance computing networks are used to improve data transfer speeds in place of archival warehouses. To facilitate this transfer and to track the location of the data, warehouse workflow managers are used. b Graphical description regarding the design and implementation of the distributed code versioning system. The master branch of the versioning system maintained by the lead bioinformatician is designed to host types of code scripts linked to scientific projects, maintaining open science standards, minimized code for publication and common code for shared project directories. The code scripts also have pre-defined code calls to access data prior to processing. This code call is designed to mimic the data hierarchy and structure of the archival data warehouse. The branch is then cloned to other members of the team, after which personnel can either begin replicating the code for their purposes or make new code scripts per requirement, which are then logged, committed and pushed to the master branch to be pulled and reproduced by other members of the team. While entire UPTIDER team is involved in the review of the code scripts, the bioinformaticians are tasked with the maintenance and upkeep of the entire code versioning system. GDPR general data protection regulation, HBM human body materials, LIMS: lab information management system, NGS next-generation sequencing, UPTIDER UZ/KU Leuven Programme for post-mortem Tissue Donation to Enhance Research. Created with biorender.com.
Developing and maintaining a distributed code versioning system
The distributed code versioning system has two key objectives: (i) ensuring the robustness of the 3 pillars to maintain research quality, and (ii) facilitating methodical downstream analyses of our data. We hosted an open-sourced system for the entire laboratory, allowing everyone in the team to access and track code6,38. The repository was structured around 4 elements to host both project specific code scripts, often maintained by one individual, and common scripts, maintained by multiple users that could easily be reused in various projects: (i) scripts for each scientific project directory, (ii) scripts and functions for QCs within the OSE, (iii) minimal code scripts that reproduced study results and (iv) scripts within shared directories for statistical analyses and data visualisation (Fig. 6b). Here, a team of 4 bioinformaticians are involved in active development and maintenance of the system which is then cloned by the entire team. The code calls for data access follow the data file structure of the data warehouses, ensuring that the appropriate files are accessed by the code scripts for every team member on their systems (Fig. 6b). Bimonthly roundtable sessions are organised, allowing code review, continuous improvements, and compliance to the system. R scripts were developed to support various sub-studies within UPTIDER, ranging from histopathological characterization, upstream and downstream processing of sequencing data to the processing of public data from cancer data repositories for our meta-analyses26. To analyse data from multiple sources, more than 30 interoperable scripts were also created to facilitate data processing and analysis. Altogether, the repository hosts all team-generated scripts, minimizing the risk of code loss. It facilitates code reviews, ensures knowledge is capitalized on, and makes the source code replicable and adaptable to support our research objectives.
Performing data curation
Data generated from UPTIDER is extensive (Fig. 1, Supplementary Fig. S1), requiring continuous efforts to ensure data consistency and accuracy. To facilitate this, the team regularly engages in eCRF and LIMS days for collective data curation of clinical and sample data and metadata. This effort is assisted by more than 30 automated QC, implemented as R scripts (Fig. 6b), on eCRF and LIMS data APIs) (Figs. 4, 5) which identifies metadata features that are either missing or incorrect according to pre-established QC rules (Supplementary Table S6). These rules co-exist with the previously mentioned internal QCs of the eCRF and LIMS. In this manner, we ensure our data and metadata are always accurate (Figs. 4, 5).
Enabling internal data reporting
To complement data curation and downstream analyses, we developed R Shiny apps39 that use the API export of the eCRF and the LIMS (Figs. 4, 5, Supplementary Fig. S4, S5). These apps helped visualize the status of clinical and sample metadata features in a user-friendly and interactive web environment. For instance, the eCRF dashboards enabled the compilation of essential clinical features by querying all patient records for metadata, including molecular subtype of the primary disease, treatments received during patient life, imaging modalities and metastatic lesions detected at various tissue sites (Supplementary Fig. S4). Similarly, the LIMS tool, which relies on complex queries, enabled the visualisation of the number of mirrored samples generated per patient across different sample conditions and timepoints (Supplementary Fig. S5). These tools help data curation and allow data exploration to develop research hypotheses and collaborations.
Training of the team
Maintaining the integrity of an OSE requires active effort, communication and training2,5. This is especially true for UPTIDER, where objectives keep evolving, and where personnel from multiple scientific areas of expertise need to interact with the environment. In this regard, regular sessions are organised by bioinformaticians to train the team in the latest changes to the OSE and to document new requirements from the team. For instance, in the eCRF, new features were added after the team was trained to document information regarding the cause of death. Considering the rapidly evolving field of liquid biopsies and biomarker detection, new protocols were developed, and metadata structures were adapted accordingly to improve annotation and complement protocol requirements. For the code versioning system, training sessions were organized to introduce new personnel to the code repository and to the scripts they will need to access. For data inventory, trainings were conducted to optimally capture the required metadata for efficient bio-tracking. These training sessions ensure accountability in data collection and reporting, foster confident engagement, which in turn improve the integrity of the OSE.
Performing data minimization and deposit
After the research manuscript has been peer-reviewed and accepted for publication, data is shared (Figs. 2, 3). Here, data and code minimization are performed to comply with the FAIR principles and GDPR, meaning only the essential parts necessary to reproduce the results are shared. In case of sensitive clinical data concerning patient information, a synthetic version of the minimised dataset is provided to prevent patient identification. The creation of interoperable code scripts within our code repository simplified code minimization. These scripts are also tested in a simulated environment on the code repository (Fig. 6b) that mimicked a code capsule environment where only minimal datasets are made available. Multiple local code re-runs are performed to ensure precise reproduction of code. Later, Code Ocean is used to deposit code and supporting metadata in a secured and installation-free, container-based environment12. Indeed, the environment outlines all the coding frameworks and supporting package libraries used to support the reproduction of code output reflecting the library versions mentioned in the research methods of the study. Moreover, the design and structure of the capsule make it convenient to export the data and environment for offline reproduction under the no lock-in philosophy of Code Ocean. During the submission of the research manuscript, code is also made available for peer-review prior to publishing the code capsule40. After the manuscript has been revised following peer-review, the open-access capsule is published with an accompanying DOI. The capsule also contains adequate supporting information regarding the published work and information for correspondence.
We apply an MIT license, which is a permissive license to use, modify and distribute our code with minimal restrictions41. We license the supporting data and metadata uploaded to the code capsule with a Creative Commons Attribution-NonCommerical-NoDerivs license (CC BY-NC-ND), to facilitate the utility of these datasets in an academic setting. As of date, 5 licensed minimized datasets and code capsules with an active DOI have been published32,42,43,44,45 from the UPTIDER study (Code availability), with 4 additional sub-studies under active development.
For all our studies where sequencing data is used, we upload the raw sequencing files to the European Genome-Phenome Archive (EGA) under protected access. EGA is a domain-specific trustworthy data repository where biomedical research data can be made publicly available46. Access to this data can be requested by external academic collaborators via a data access committee (DAC) on the EGA portal. Thereafter, following an approval from the UZ Leuven ethics committee, a data transfer agreement needs to be signed prior to accessing our sequencing data25,47. For access to samples from our biobank, following correspondence with the team, it requires approval by our ethics committee and a signed material transfer agreement25,47. These agreements ensure that the research investigation aligns with responsible secondary use of patient material and data, while also complying with ethical requirements from UZ Leuven and GDPR25. These conditions are effective only if the patient consented to share their tissue samples and derived data in their informed consent form, which was recorded in the UPTIDER eCRF at study inclusion.
OA publishing
OA publishing remains the quickest way to communicate your research to a global audience while benefiting from increased visibility, discovery and reporting16. Most OA journals are also peer-reviewed, ascertaining the work quality. We opted for gold and diamond OA publishing as they enable unlimited public access to the research work upon publication15. Our commitment to OA publishing is in line with the mandates of federal funding agencies, making our work visible, accountable and transparent. This in turn, enables better research and collaboration, improving public health outcomes. As such, all research and/or review articles containing data derived from the UPTIDER study have been published in either a gold or a diamond OA journal32,42,45,48,49,50. Furthermore, protocols used to achieve our outputs are fully detailed in our publications for peer-replication. As team contributions are well documented in our publications, respective team members remain available for further correspondence.
Dissemination and communication
Sharing the experience of building and maintaining an OSE to yield high-quality research outputs could provide a much-needed incentive for other researchers to develop their own. As open science principles are gaining traction, calls for implementation at the institutional level are increasing4,13. To this end, we presented our OSE internally, at open science conferences and cancer congresses to raise awareness on the benefits and the advantages of investing in an OSE for translational research51,52. We also advised and guided other labs that were interested in customizing an OSE to their research requirements. Expanding this, we also provide a list of software and infrastructural components used within our OSE to help researchers identify the appropriate tools for their own use (Supplementary Table S7).
Designing a data management plan
For effective research, a DMP is generally mandatory at the project level. It provides the necessary information concerning the acquisition, consumption, storage and re-use of research data to support translational research objectives, while complying with ethical and regulatory frameworks. A DMP also contains information on personnel responsible for correspondence of study progress and the expected costs for data processing and study publication. Here, prior to developing project-specific DMPs and designing the components of the UPTIDER OSE, we designed an overarching DMP at the laboratory level, to be compatible with open science, GDPR and FAIR principles while promoting sustainable translational research in metastatic breast cancer. Building an overarching DMP helped keep a global overview of the data workflow. It also contributed to proper environment design and the generation of the mandatory project-specific DMPs. More importantly, the DMPs are active and dynamic documents that keep evolving with the growing needs of each research study.
Assessment and continuous improvement
To sustain an evolving OSE, it is important to implement (self)-assessment strategies to identify strengths and weaknesses. We leveraged our institution’s commitment to high-quality RDM services, including a self-assessment methodology based on published models53,54. This tool evaluates the OSE in terms of research planning, data collection, processing, and storage. Based on the assessment outcomes, RDM support staff provide guidance and feedback. We participated in the pilot phase of the RDM assessment, evaluating our OSE’s compatibility with open science principles (Supplementary Fig. S6). After the session, we implemented improvement suggestions and scheduled yearly assessments for continuous improvement, ensuring our environment remains compatible with open science standards and evolves with the UPTIDER project needs.

